Quality control report
Rastair comes with a helper script, written in R, which generates a comprehense QC report for your library.
If not using the docker image: The QC tool requires a working installation of R with RMarkdown, data.table, ggplot2 and argparser libraries. You can install all of them at once with
install.packages(c("argparser", "data.table", "ggplot2", "rmarkdown"))
For V-bias and GC-bias plots, it also requires Rsamtools from Bioconductor. This can be installed with
if (!require("BiocManager", quietly = TRUE)) install.packages("BiocManager") BiocManager::install("Rsamtools")
It also requires pandoc for rendering and bcftools for GC and CpG bias plots.
Generating QC reports
The qc report can be generated directly from a bam file, or from pre-generated per-read and per-position bed files:
mkdir -p test_qc
rastair mbias --output-prefix test_qc --bam test.bam
This will produce a file called qc_report.html in the test_qc directory.
You can selectively disable the V-bias and GC/CpG bias plots to speed up processing and/or reduce dependencies:
mkdir -p test_qc
rastair mbias --output-prefix test_qc --no-vbias --no-gc --bam test.bam
Elements of the QC report
Conversion of methyl-C to T should be independent of where in a sequencing read the methylation ocurred. However, in reality, a number of experimental procedures can affect the evenness of conversion. For example, sonication or enzymatic digestion of genomic DNA can produce single-stranded ends. When these are repaired for A-tailing, the newly synthesized parts will be unmethylated.
To identify any potential issues in your data, we are providing two types of plots to investigate the evenness of methylation:
M-bias
The classic "methylation bias" (aka M-bias) plot shows the average methylation per position in a read:
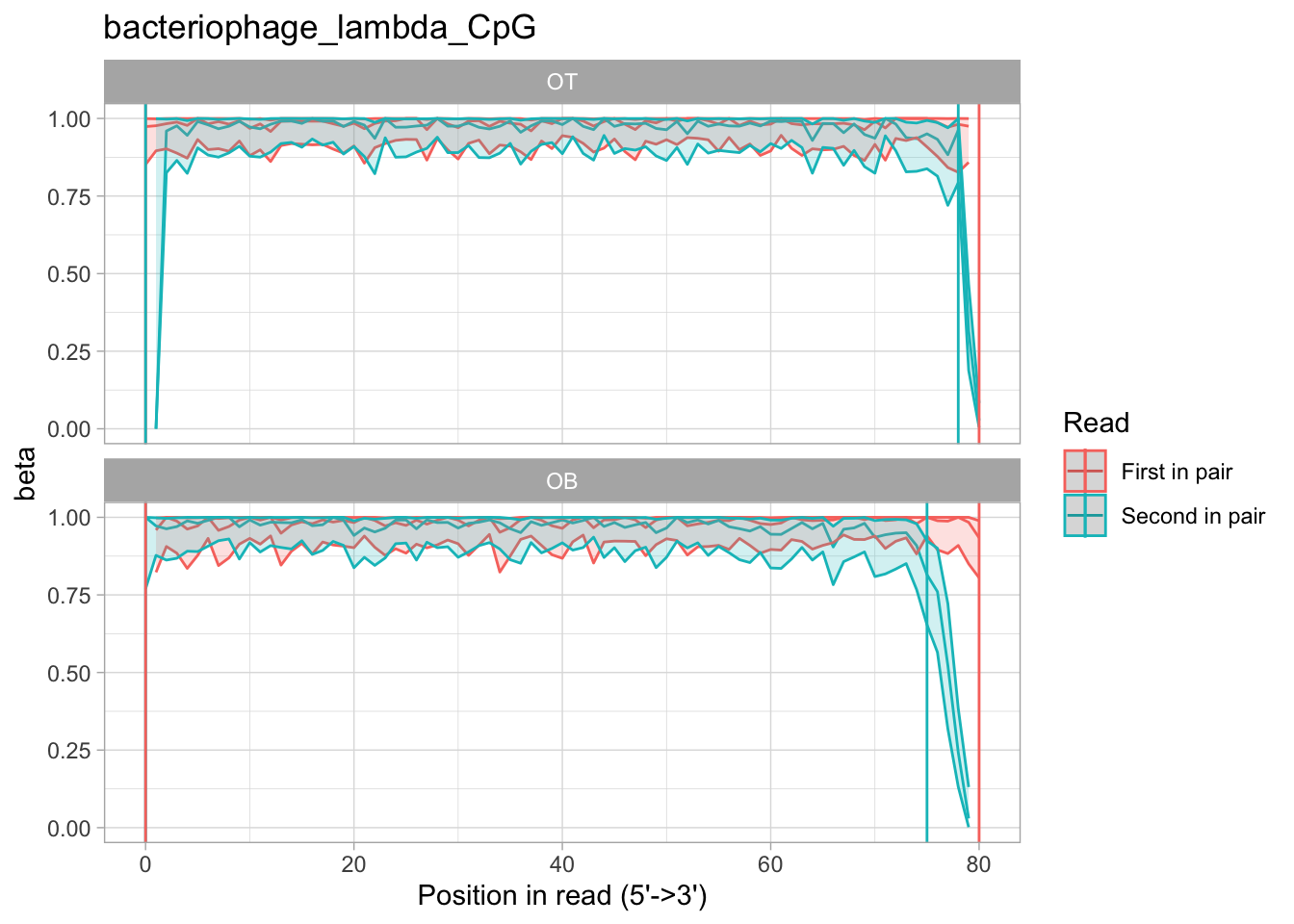
We distinguish between OT and OB fragments, and show average methylation for R1 and R2 reads separately. This is an example of an enzymatically fully methylated piece of DNA that we use as a control, so methylation is nearly 100% across most reads. However, you can see that the ends of R2 of OB fragments seem to have lost methylation information. This could be corrected with the --nOT and --nOB arguments to rastair call.
To facilitate choosing the right cutoff parameters, we also generate a text file for each processed chromosome that contain the estimated optimal --nOT and --nOB settings, respectively.
In this example, the script produced this:
cat bacteriophage_lambda_CpG_cutoffs.txt
0,0,0,2
0,0,0,5
In other word, it suggested we set --nOT 0,0,0,2 and --nOB 0,0,0,5, ie remove the last 2 and 5 bases of R2 in OT and OB, respectively.
V-bias
In some cases, for example when DNA is enzymatically digested in vivo or in vitro, there might be a wide range of fragment sizes. We have observed in some instances that fragment size has an impact on methylation bias per read position. To visualise this, we provide what we call a "V-bias" plot:
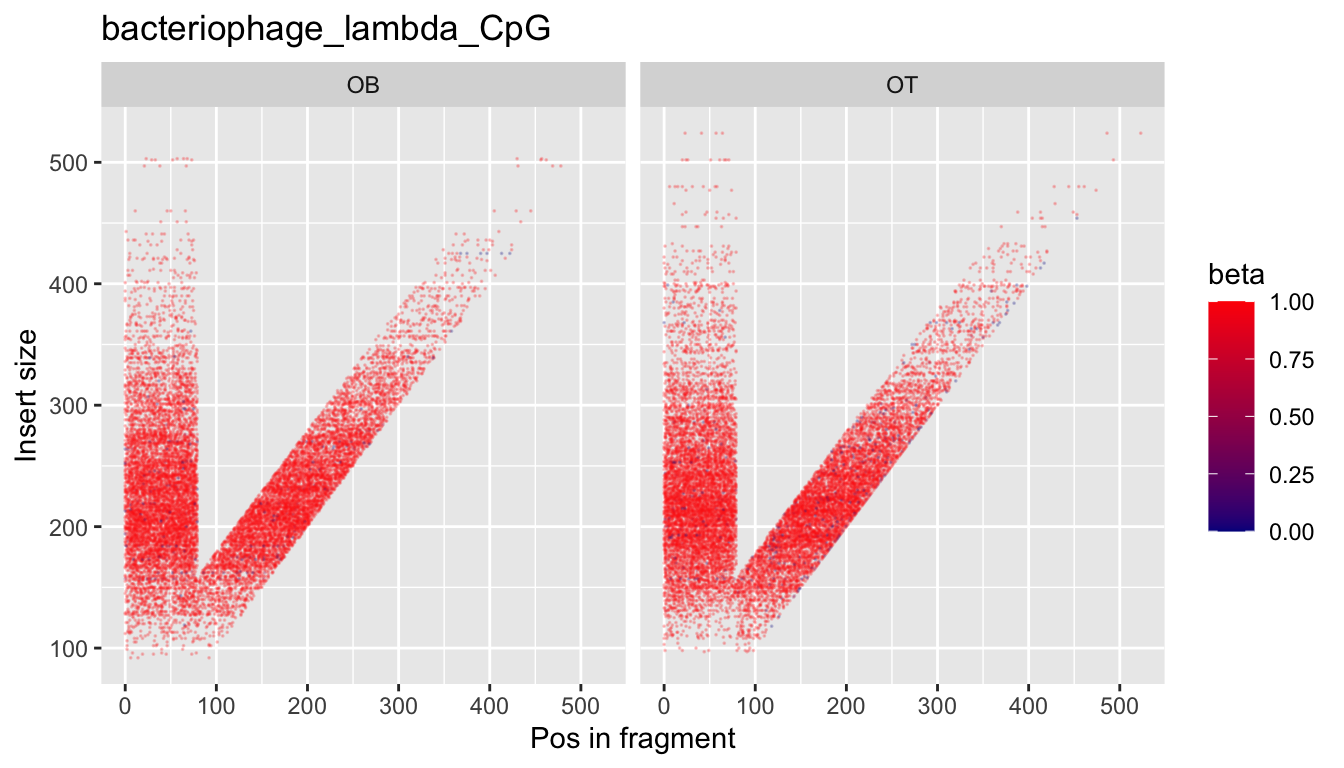
Here, colour denotes methylation, while the y-axis reflects the length of the original fragment, assuming paired-end sequencing.
GC/CpG bias
We also provide some plots to visualise the relationship between local GC content and CpG density per read vs methylation and coverage:
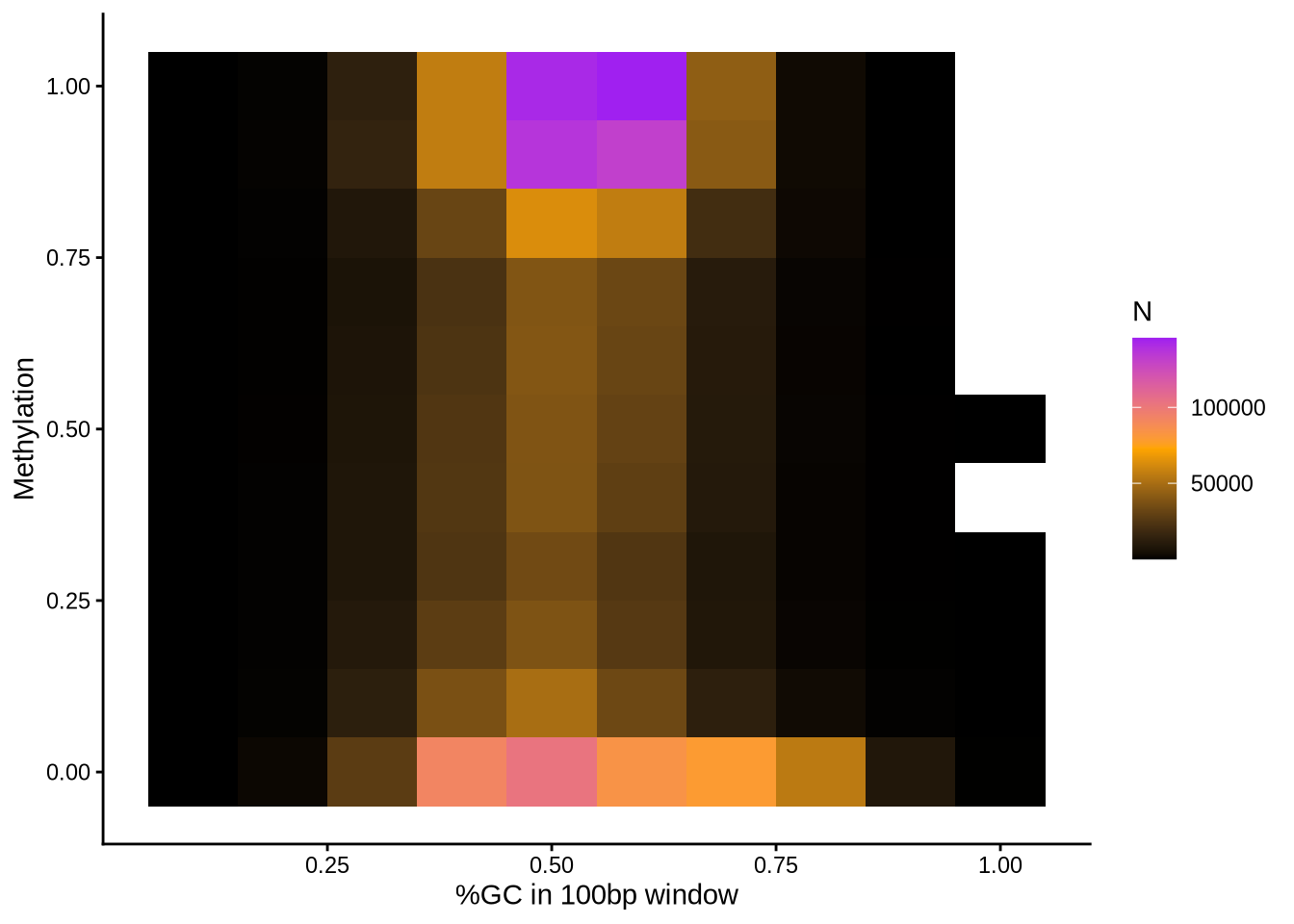
Here, you can see whether highly/lowly methylated reads fall into high/low GC-content regions.
Another occasional issue is coverage differences depending on CpG density of the read:
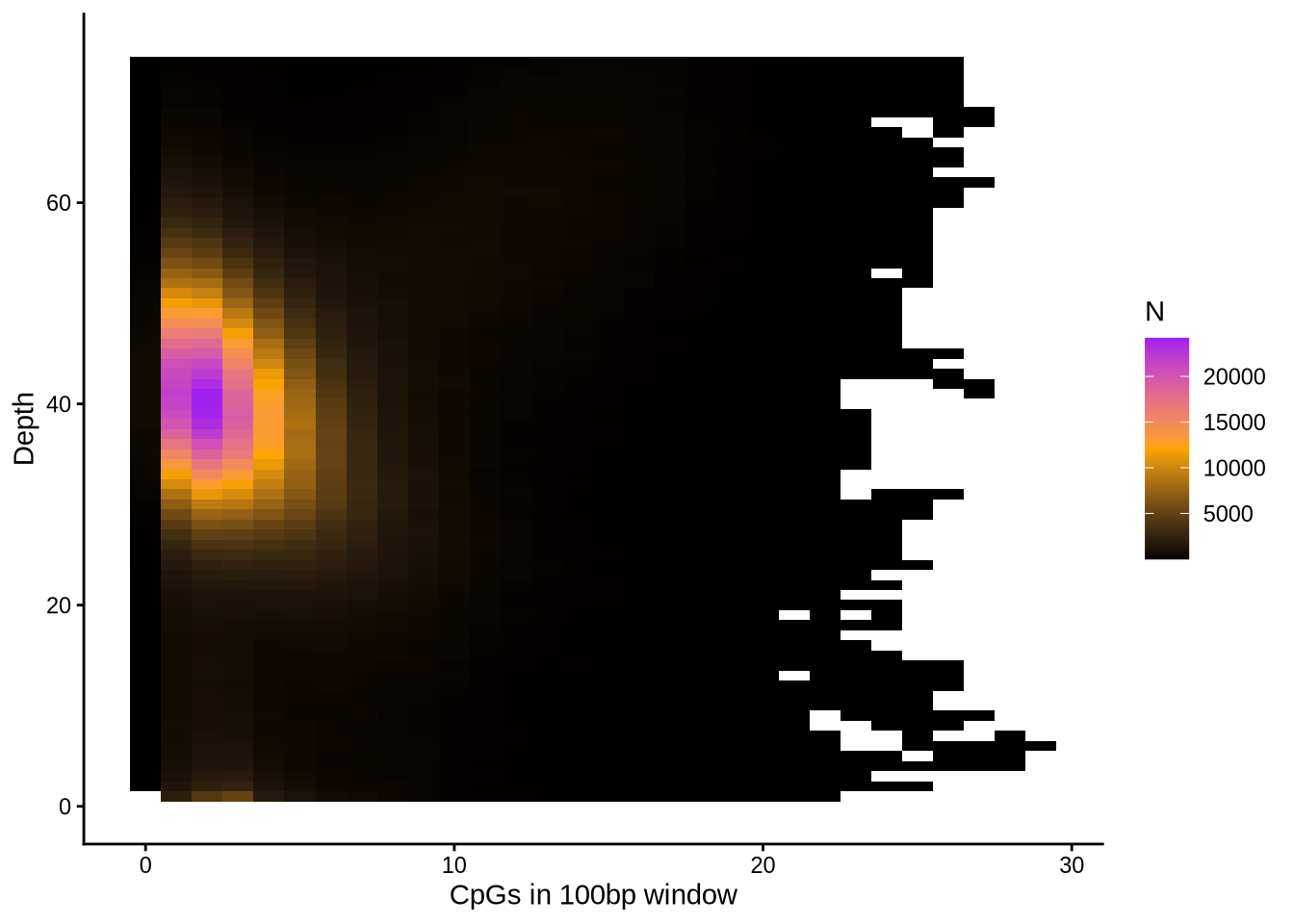
Here, you can see that there's a slight negative correlation for 100bp regions with many CpGs to have lower coverage.